Every MLPA experiment must include at least three independent DNA reference samples. For most digitalMLPA experiments it is also recommended to use at least three independent reference samples. Reference samples should be derived from healthy individuals that are not expected to have any copy number changes or mutations in your region of interest. When using more than 21 samples, one additional reference sample should be added for every seven samples.
A no-DNA control is recommended for MLPA experiments, and is optional for digitalMLPA experiments. Inclusion of one or more positive control samples, with a known deletion, duplication, point-mutation and/or methylation status, is recommended when possible as an aid in result interpretation. In some cases it may also be useful to include a reaction with commercial DNA for troubleshooting purposes.
Reference samples
MLPA and digitalMLPA are relative techniques based on the analysis of relative changes in probe signals or probe read counts, respectively. A single sample cannot provide the information needed to estimate copy number changes without something to compare to. Reference samples are samples where the target sequences of interest are assumed to have a "normal" copy number and, in the case of MS-MLPA, a "normal" methylation status. Usually, reference samples are DNA samples obtained from healthy individuals.
It is strongly recommended to use reference samples that have been purified using the same method, and that have been derived from the same type of tissue as the test samples. This minimizes non-biological differences between test and reference samples and minimizes variation. Multiple reference samples are needed to estimate the reproducibility of each probe. Inclusion of multiple reference samples also helps prevent false positive/negative results due to experimental issues or biological variation in the reference samples/reactions.
Reference samples should be distributed “randomly” over the experiment to avoid bias. The image below shows an example.
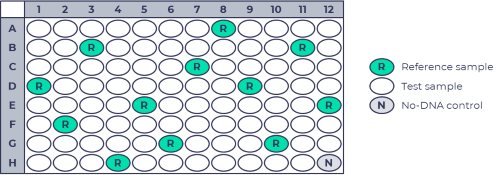
Reference samples are required in most digitalMLPA experiments. In some cases, reference samples can be omitted when enough independent samples (from different families) are run at the same time, and when the chance for each individual probe being deleted or duplicated in a sample is very low. If no dedicated reference samples are used, all samples should be specified as test samples in the data analysis in Coffalyser digitalMLPA. Outliers with copy number variations will automatically be ignored as the data analysis makes use of medians instead of averages. Experiments with some digitalMLPA probemixes may more readily meet these requirements. The number of samples that should be included depends on the application and the experimental setup, but in general a larger number of samples will be better (more information). However, inclusion of reference samples is never a bad idea, as they can also provide information in case results are not as expected. If a digitalMLPA probemix contains X- or Y-chromosome target probes, sufficient reference samples should be present of each sex.
When analysing tumour samples, reference samples should still be treated as similarly as possible. Ideally, reference samples are derived from similar, but healthy tissue, that has been treated in the same way as the test samples. This may include FFPE-treatment (see this article for more details). For the detection of methylation changes in tumour samples, it is important to realize that methylation is cell-type specific, and can vary between tissues.
No-DNA controls
A no-DNA control is a reaction in which TE0.1 (10 mM Tris-HCl pH 8.0 + 0.1 mM EDTA) is used in place of the sample DNA.
Inclusion of a no-DNA control in every MLPA experiment is strongly recommended. The results can be used to check for contamination of reagents or equipment, and for the formation of non-specific peaks. More information on how to interpret an MLPA no-DNA reaction can be found in this article.
Inclusion of a no-DNA control in a digitalMLPA experiment is not essential. The digitalMLPA technique has been optimised to ensure that almost all reads from non-specific amplicons are removed during data analysis. In addition, other quality control measures can be used to check for contamination. The SNP-specific control probes included in each digitalMLPA probemix are used to check for contamination of the sample DNA or reagents with DNA from another individual. The absence of a substantial number of reads from barcodes not in use in the experiment can be used to check for contamination with reads from a previous experiment.
Positive control samples
Positive control samples can be used to check whether the entire procedure, including data analysis, performs as expected. While positive samples can be useful, they are not required.
Like reference samples, positive control samples are preferably purified using the same method and from the same type of tissue as the test samples. If no positive control samples are available, suitable samples can sometimes be ordered from an online biorepository. MRC Holland has very limited access to patient samples and cannot supply positive samples. This article lists a large number of commercially available samples that have been tested by MRC Holland with our products.
Commercial DNA
If you experience recurring issues with the quality of your reactions, it can be useful to include a reaction with DNA that is known to be of high purity and quality, such as commercially available DNA. We recommend using Promega Human Genomic DNA (catalog number G1471 for male DNA or G1521 for female DNA).
If reactions with this commercial DNA work well but reactions on your own samples have issues, there may be a disruptive impurity present in your samples (read more about sample purity and quality). If reactions with this commercial DNA show the same issues as reactions on your own samples, there is an increased chance that the issues are caused by experimental factors.
Commercial DNA should never be used as reference sample, as it has not been treated in the same way as your own samples (see the section on reference samples above). Routine inclusion of reactions with commercial DNA is not recommended, as it is generally unnecessary. Inclusion of a reaction on commercial DNA might also yield no extra information if you have positive samples or reference samples that are known to work well with the probemix in question, or if your experiment includes one of our sample DNAs.
If you experience reaction issues that require troubleshooting, you are also welcome to contact us for troubleshooting assistance.